The Swiss Ageing Citizen Reference (SACR)
SACR is a driver project of the Swiss Personalized Health Network (SPHN) aimed to:
- Harmonize and apply data and biomaterial from existing Swiss citizen cohorts to build the well-embedded public health pillar of personalized health in Switzerland.
- Grant personalized health researchers responsible access to prospectively obtained information on health and health risks, to geocoded residential and work address history, and to archived biomaterial for testing the clinical and public health utility of candidate biomarkers and exposome features in predicting aging-related intermediate biomarkers (DNA methylation; MRI derived brain features), and subsequently aging-related (multi-) morbidities.
SACR puts together three population-based cohorts (SAPALDIA, CoLaus/PsyCoLaus, and SKIPOGH) to create a novel data set of sample size >900, including brain MRI, DNA methylome of prospective biospecimen, and aging phenotypes. The SACR database will make a unique contribution to the aging research in itself or by contributing healthy reference data for case-control analysis.
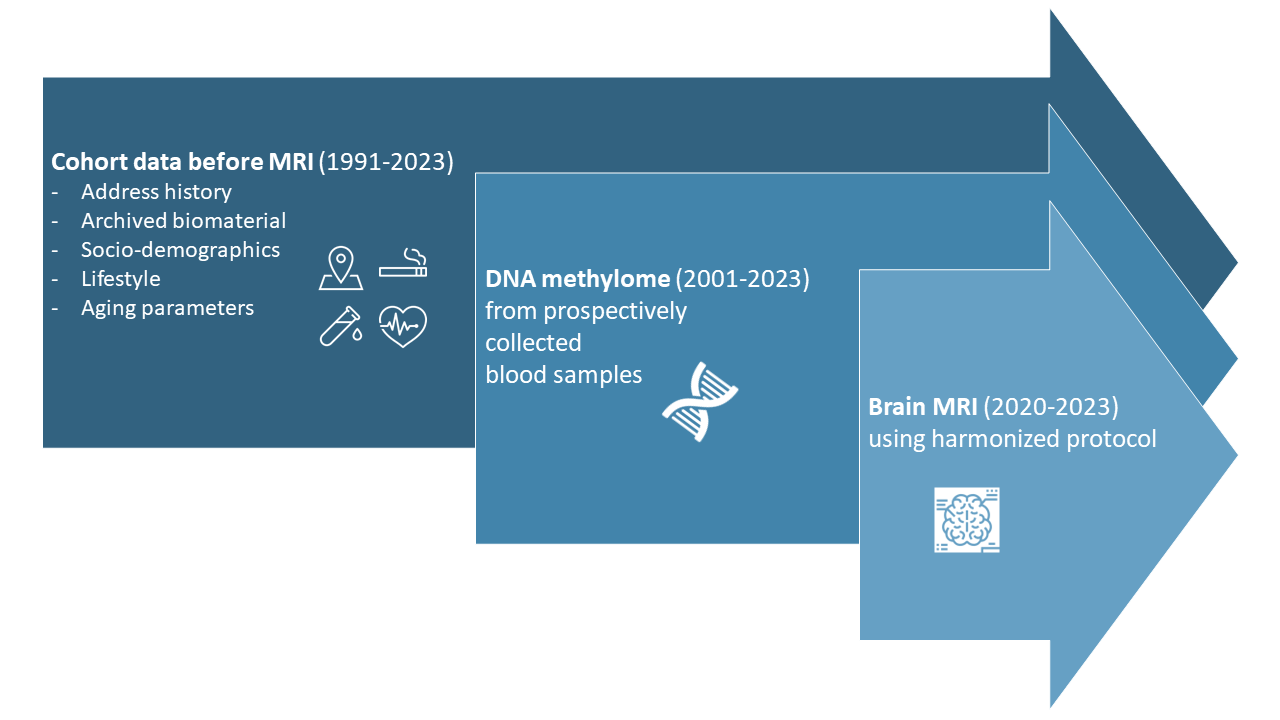
SACR database
Deep phenotyping in the context of previous cohort examinations
The SACR standard dataset was developed by matching variables from the population-based cohorts SAPALDIA, CoLaus/PsyCoLaus, and SKIPOGH. Variables comprising personal characteristics, lifestyle and adiposity, and healthy aging related phenotypes have been assembled and harmonized across the three cohorts.
Brain MRI features
902 participants of SACR have had their MRI taken at one of the SACR study centers. The image processing pipeline is similar to the one described by Trofimova et al. (Commun Biol. 2023 Apr 10;6(1):392) and transforms the raw voxel-based data into segmented data where standardized percentage intensity values are given per anatomical region and modality. The anatomical region definitions are a custom selection of mesh terms, UMLS terms and anatomical regions defined in the neuromorphometrics challenge.
DNA methylome
Blood samples mostly prospectively sampled from 891 out of the 902 participants with MRI have been identified. DNA methylome analysis is performed using the Illumina BeadChip Array (v2.0). Bisulfite treatment and methylome assessment are conducted on DNA samples in a randomized order, which minimizes batch effects. The randomization scheme has been described in detail in this document. The DNA methylome data will be processed following the same principle as the previous DNA methylome analyses in SAPALDIA (Jeong et al 2022; Jeong et al 2019), including background and dye-bias correction, beta-mixture quantile normalization, and batch effect control.
Codebooks
Note that the SACR cohorts, which have been implemented with different objectives, have much richer information over longer time period, while the SACR codebooks contain a more limited, but fully harmonized number of variables. The whole spectrum of information can be explored via meta-data databases:
How to get access to the SACR database?
Interested researchers can fill in the SACR research application form and submit it to nicole.probst@swisstph.ch. The SACR consortium board will review the application and inform the applicant. Upon the acceptance of the application, project-specific DTA/MDTA will be prepared by SACR consortium. Costs for data access and analysis capacity may arise for the applicant.





