Tuberculosis Ecology and Evolution
Tuberculosis (TB) is among the ten top causes of human death and remains the leading cause of mortality from a single infectious agent globally. TB is caused by a group of bacteria known as the Mycobacterium tuberculosis Complex (MTBC), yet we are only beginning to understand the genetic diversity within the MTBC and how this diversity shapes TB transmission, drug resistance, and patient treatment outcomes. To address these challenges, our group studies the causes and consequences of this diversity, investigating how evolutionary forces drive bacterial adaptation and influence the epidemiology of TB across the world.
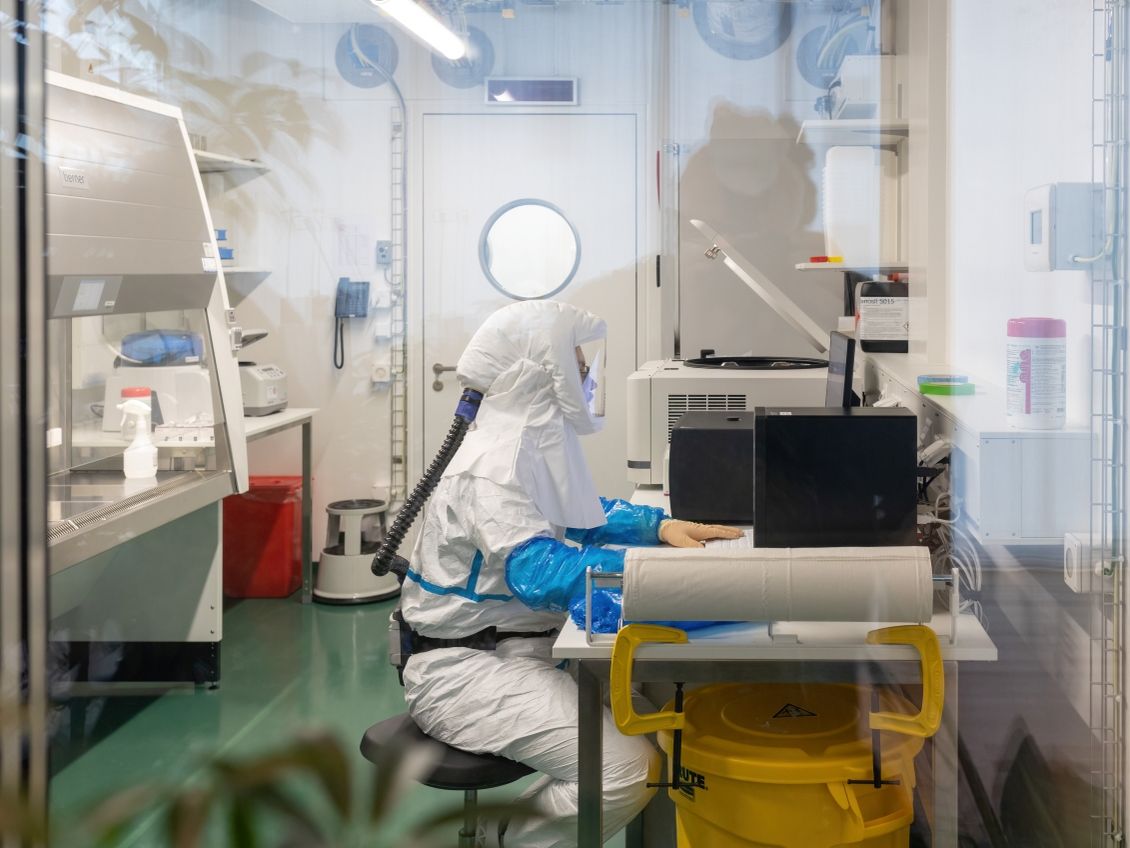
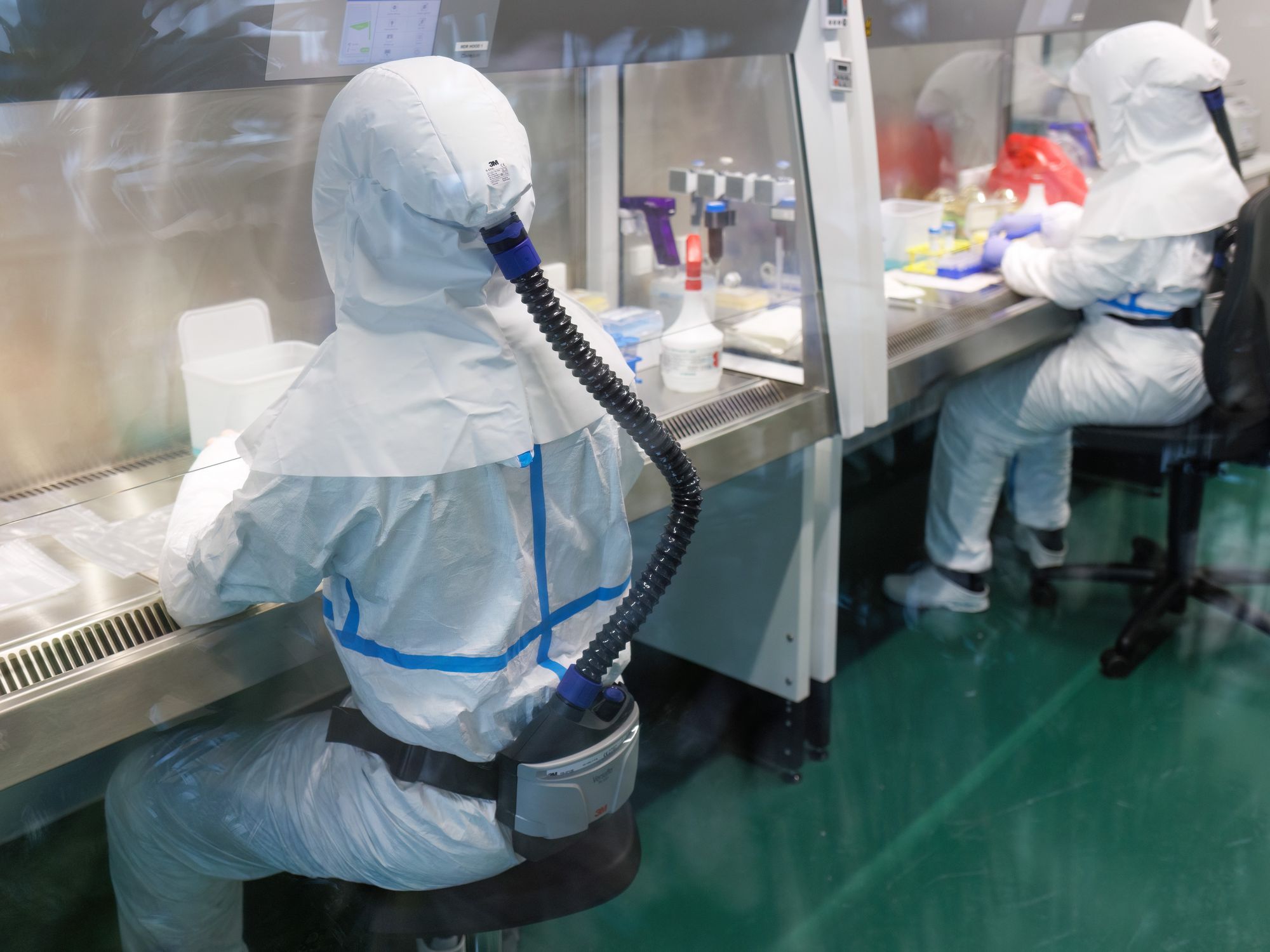
Through state-of-the-art BSL3 facilities, expert genomics capacity, and international collaborations, we investigate the fundamental biology of the MTBC, alongside the molecular epidemiology of TB in Switzerland and globally. Our work spans from understanding the genetic population structure of the MTBC to tracking transmission dynamics in real-time, revealing how evolution operates within individual patients and across patient populations.
Our research focus
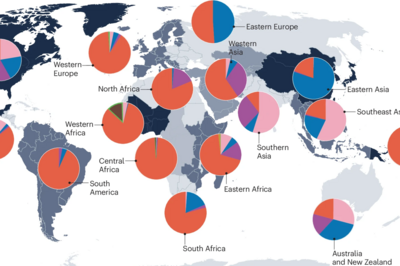
The global population structure of the MTBC
The MTBC comprises ten human-adapted phylogenetic lineages associated with different geographical regions, and several lineages adapted to different wild and domestics animals. We use comparative whole genome sequencing to study the differences between these lineages and the evolutionary forces shaping their genomic diversity. By combining various omics technologies with functional assays and epidemiological data, we investigate the phenotypic consequences of this diversity – revealing how evolutionary history influences host specificity, virulence, transmissibility, and drug susceptibility. Read more
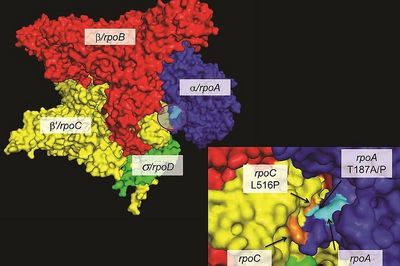
Ecology and evolution of drug-resistant MTBC
Drug resistance threatens global TB control efforts. When drug-resistant bacteria first emerge, they often transmit poorly due to fitness costs associated with drug resistance mutations. However, evolution is continuous and drug-resistant strains readily adapt through compensatory mutations that restore transmissibility. We study these evolutionary processes to understand how highly transmissible drug-resistant strains emerge and spread, thereby informing strategies to better prevent and control antimicrobial resistance. Read more
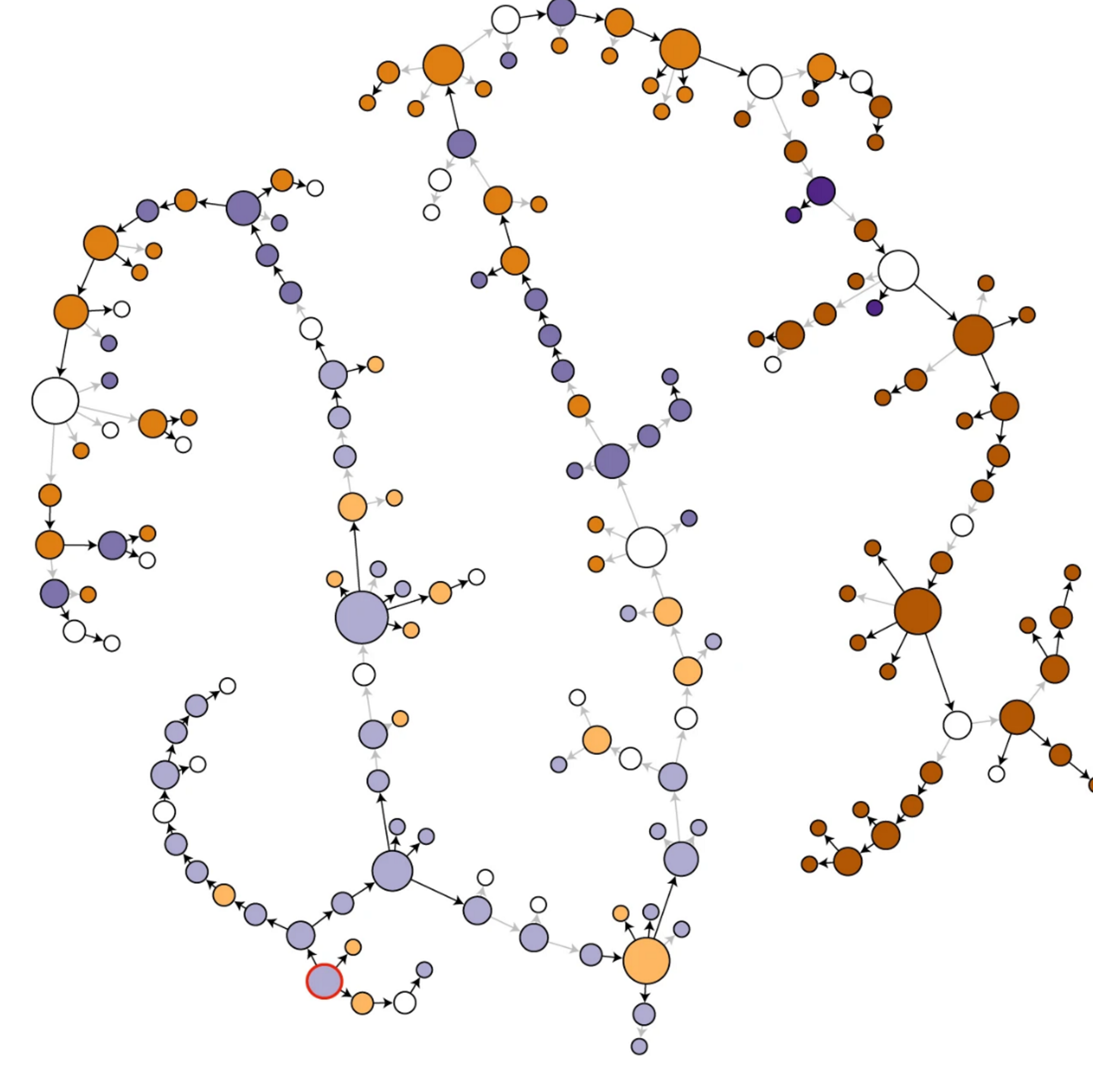
Genomic epidemiology of TB
Whole-genome sequencing has revolutionized how we track TB transmission. We use genomic epidemiological approaches to study TB transmission dynamics in Switzerland and TB-endemic countries, reconstructing transmission chains and identifying outbreaks. We also explore the micro-evolution of MTBC within individual patients during treatment, revealing how bacteria adapt to therapeutic pressure in real time. Read more
Ongoing projects
Historical contingencies in extensively drug-resistant TB evolution
In Georgia, 30% of MDR-TB cases are linked to incarceration. This project explores how prisons facilitated the emergence of highly transmissible MDR-outbreak strains and how their evolutionary histories influence adaptation to new treatment regimens including bedaquiline and pretomanid. Using over 30,000 clinical isolates spanning three decades, we combine long-read sequencing and proteomics to understand historical contingencies in bacterial adaptation. Read more
MTBC evolution across timescales and drug resistance development
In collaboration with TB Alliance, we study the MTBC strain collection from clinical trials that led to approval of bedaquiline- and pretomanid-containing regimens by applying long-read sequencing and comparative proteomics to study genome evolution, drug tolerance, and resistance emergence across different evolutionary timescales. Building on these ‘real-world experiments’, we study how bacterial variation influences patient treatment outcomes and resistance development to new anti-TB drugs. Read more
Linking within-host and between-host evolution
The ECOEVODRTB project defines genomic characteristics and evolutionary forces shaping MDR MTBC populations within individual patients over time and across body compartments, comparing within-patient evolution to between-patient transmission dynamics. Combining population genomics and genomic epidemiology with laboratory experiments, we reveal how suboptimal treatment and phenotypic drug tolerance drive resistance evolution inside patients – and how this connects to transmission between patients. Read more
Asymptomatic TB and the virulence-transmission trade-off
Why do "ancient" MTBC lineages persist despite lower virulence than "modern" lineages? We hypothesize they pursue a different evolutionary strategy – causing asymptomatic TB that extends infectious periods, maximizing transmission over time. Working in collaboration with the Ifakara Health Institute (IHI) in Tanzania, we compare genomic, phenotypic, and epidemiological characteristics of asymptomatic versus active TB across MTBC lineages to test for the trade-off between virulence and transmission. Read more
PANGenS – Pan-Africa genomic surveillance network
Led by the University of Ghana with Swiss TPH partnership, this network strengthens Africa's capacity for the genomic epidemiology of TB, malaria, and emerging pathogens. The project establishes collaborative frameworks spanning wet lab to bioinformatics training, develops biobanking and data management infrastructure, and implements proof-of-concept surveillance studies – equipping African scientists with cutting-edge skills while enhancing disease surveillance across the continent. Read more



Collaborations and networks
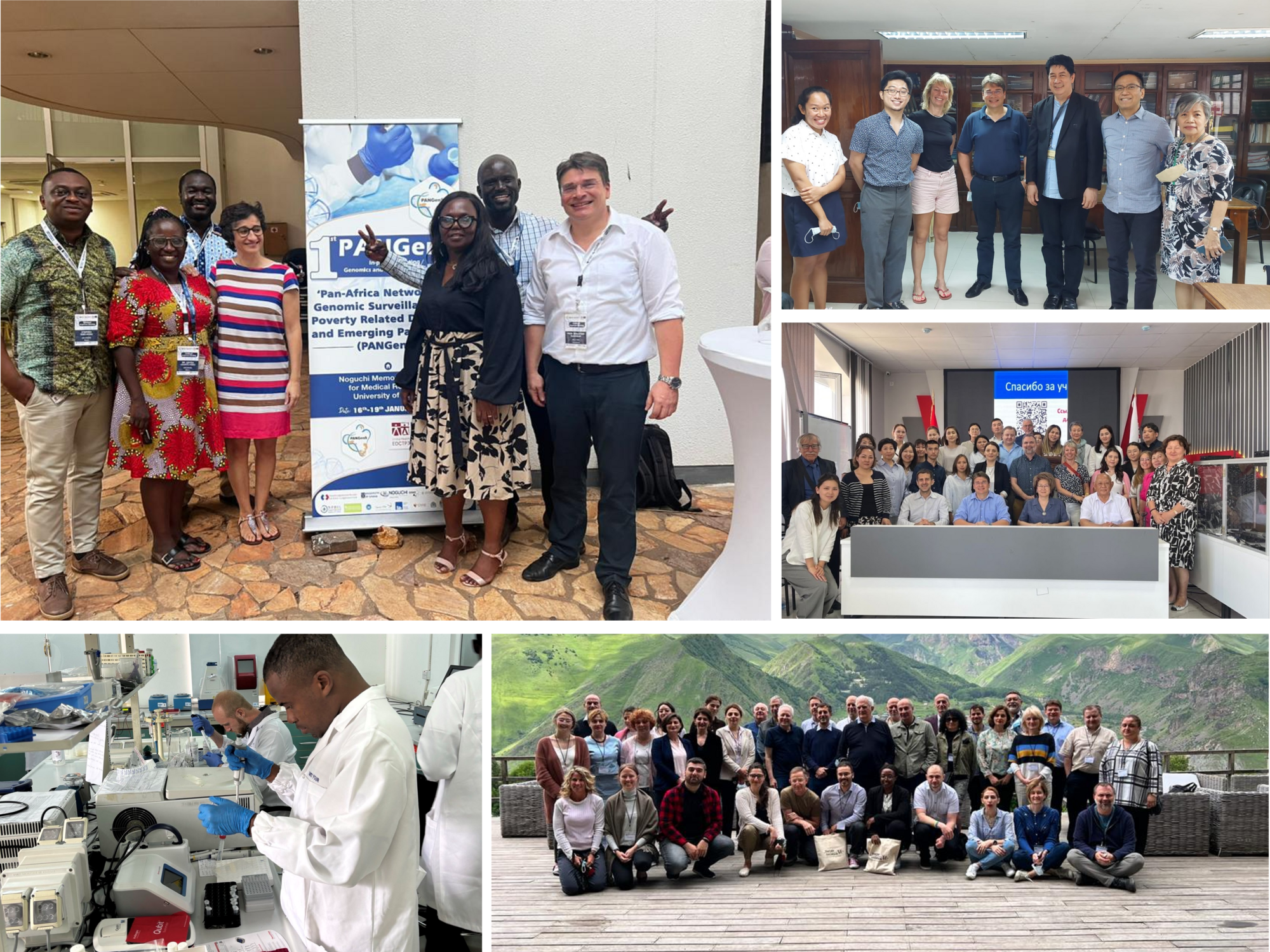
Our research relies on partnerships with collaborators in TB-endemic countries, fostering multinational cooperation, capacity strengthening, and access to critical clinical and demographic insights. Our recent collaborative partners include institutions in Ghana, the Philippines, Tanzania, Georgia, and Kyrgyzstan. Our key long-term partnerships include the National Centre for Tuberculosis and Lung Diseases in Georgia, the Ifakara Health Institute in Tanzania, and the Noguchi Memorial Institute for Medical Research in Ghana with all of whom we have been collaborating for more than a decade.
Selected publications
related to our work on the ecology and evolution of drug resistance in the MTBC:
related to our work on the general ecology and evolution of the MTBC:
Frontiers | A New Phylogenetic Framework for the Animal-Adapted Mycobacterium tuberculosis Complex
Local adaptation in populations of Mycobacterium... | F1000Research

 Amir Banaei Esfahani
Amir Banaei Esfahani
 Daniela Brites
Daniela Brites
 Sevda Kalkan
Sevda Kalkan
 Chloé Marie Loiseau
Chloé Marie Loiseau
 Veronica Misana
Veronica Misana
 Gian Schüpbach
Gian Schüpbach