Unit | Tuberculosis Ecology and Evolution
Tuberculosis is an infectious disease caused by the bacterium Mycobacterium tuberculosis (Mtb), one of the deadliest pathogens known to humankind. In the Tuberculosis Ecology and Evolution Unit, we study the causes and consequences of genetic diversity in the Mycobacterium tuberculosis complex (MTBC).
Through the combined strengths of state-of-the-art BSL3 facilities, expert genomics capacity, and international collaborations, we’re equipped to investigate the fundamental biology of Mtb alongside the molecular epidemiology of TB across the world.
We focus on understanding the global diversity of the MTBC, the evolutionary forces that drive this diversity, and the phenotypic consequences of this diversity for the biology and the epidemiology of TB.
Partnerships
Our work relies on our long-term partnerships with collaborators in TB-endemic countries, fostering multinational cooperation, capacity building while gaining demographic and clinical insights. Recent partnerships include collaborations with entities in Ghana, the Philippines, Tanzania, Georgia and Kyrgyzstan.
Main Research Areas of the Tuberculosis Ecology and Evolution Unit
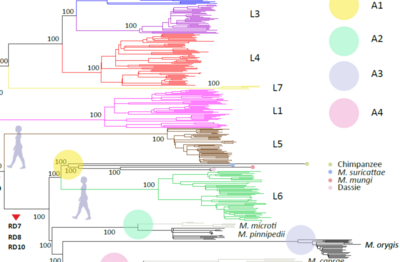
The Global Population Structure of the MTBC
The human-adapted MTBC comprises seven phylogenetic lineages that are associated with different regions of the world. We use comparative whole genome sequencing to study the differences between these lineages and the evolutionary forces shaping this diversity. We combine various –omics technologies with functional assays and epidemiological data to investigate the phenotypic consequences of this diversity. Read more
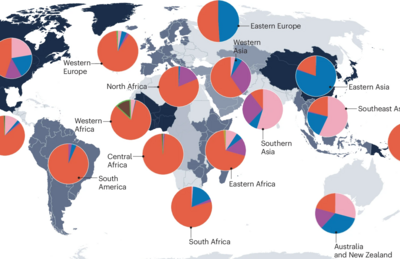
Genomic Epidemiology of TB
Recent advances in whole-genome sequencing have revolutionized molecular epidemiological investigation of TB. We use such genomic epidemiological approaches to study the transmission dynamics of TB in Switzerland and in TB-endemic countries. We also explore the micro-evolution of MTBC in individual patients during treatment. Read more
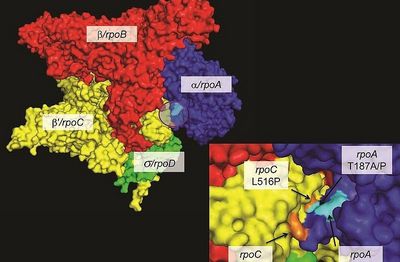
Ecology and Evolution of Drug-Resistant MTBC
Drug resistance poses a growing threat to global health. When drug-resistant bacteria first emerge, they are often less transmissible than susceptible strains – this is because drug resistance in bacteria is often associated with a reduction in Darwinian fitness. However, evolution is a continuous process, and drug-resistant bacteria readily adapt and regain the ability to transmit. This process is mediated by compensatory mutations. Read more

 Amir Banaei Esfahani
Amir Banaei Esfahani
 Sonia Borrell
Sonia Borrell
 Daniela Brites
Daniela Brites
 Adrian Denz
Adrian Denz
 Galo Goig
Galo Goig
 Sevda Kalkan
Sevda Kalkan
 Chloé Marie Loiseau
Chloé Marie Loiseau
 Valerie March
Valerie March
 Veronica Misana
Veronica Misana
 Gian Schüpbach
Gian Schüpbach